High-throughput genetically regulated omics imputation from chromosome-split PLINK2 genotype data.
The gromtools package enables high-throughput imputation of genetically regulated omics values from chromosome-split PLINK2 genotype data using additive linear SNP models.
Major functionality of the gromtools package:
| Function | Description |
|---|---|
read_db_dir() |
Load model SQLite databases into a weights table |
grom_impute() |
Build sparse weights and write .grom, .gid, and .sid outputs |
grom_read() |
Stream selected model, gene, and sample combinations from an existing .grom output |
Resources
- Minimal getting started vignette
Motivation
Large-scale genetically regulated omics workflows need a compact interface for loading prediction weights, streaming genotype data, and writing output in a format that can be read back efficiently without materializing the full matrix in memory. gromtools is built around that workflow. It converts model weights into a sparse binary representation, streams chromosome-split .pgen inputs, computes predicted values using additive SNP effects, and writes the result to an on-disk .grom matrix with matching .gid and .sid index files.
The package is designed around three user-facing steps: read model databases, run imputation, and read back selected results. Example synthetic inputs for the full workflow are bundled with the package.
Why GROMTools?
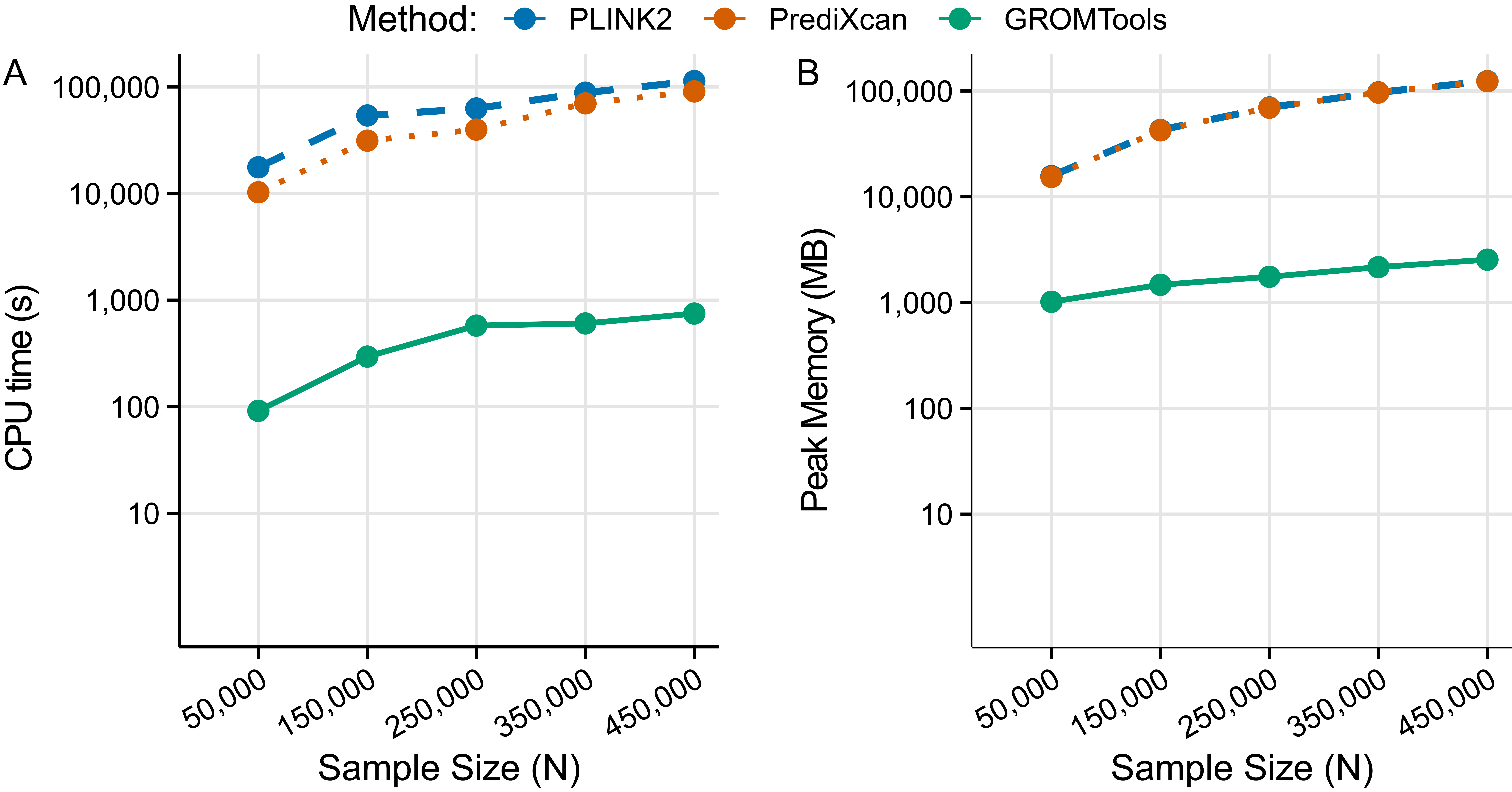
GROMTools substantially reduces the computational cost of genetically regulated gene-expression (GReX) imputation. In chromosome 1 benchmarks across 50,000 to 450,000 individuals, it consistently outperformed PLINK2 and PrediXcan in both CPU time and peak memory usage. All analyses were conducted single-threaded on the same machine to ensure a fair comparison. Across these benchmarks, GROMTools was about 69x to 209x faster and used about 15x to 49x less peak memory, depending on the comparator and sample size. This makes it particularly well suited for large-cohort and biobank-scale analyses.
Install
devtools::install_github("voloudakislab/gromtools")